Summary
Clan
This family is a member of clan (CL00129), which contains the following 3 members:
cHP DENV_SLA Flavivirus-5UTRWikipedia annotation Edit Wikipedia article
The Rfam group coordinates the annotation of Rfam families in Wikipedia. This family is described by a Wikipedia entry Flavivirus 5' UTR. More...
This page is based on a Wikipedia article. The text is available under the Creative Commons Attribution/Share-Alike License.
Sequences
Alignment
Seed alignment view
Download
Download a gzip-compressed, Stockholm-format file containing the seed alignment for this family. You may find RALEE useful when viewing sequence alignments.
Submit a new alignment
We're happy receive updated seed alignments for new or existing families. Submit your new alignment and we'll take a look.
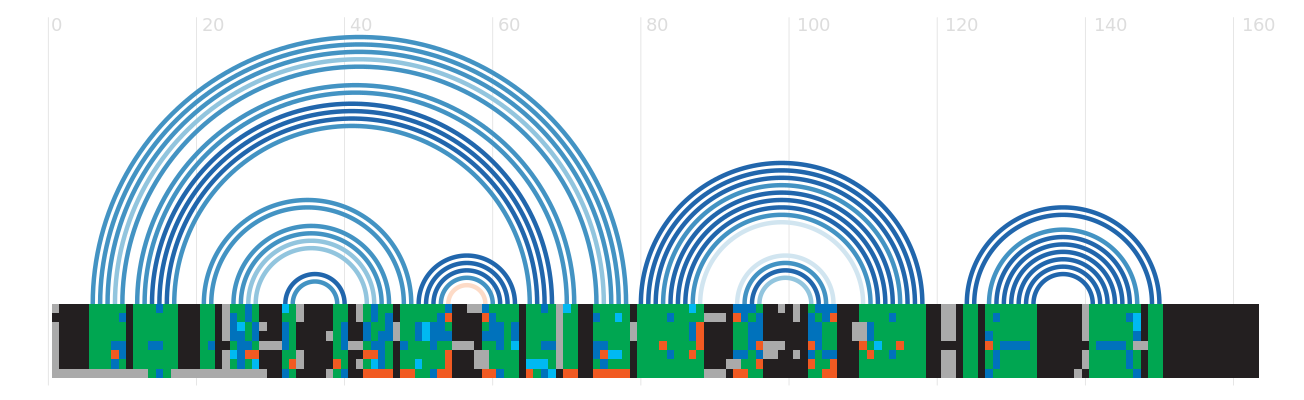
Secondary structure
Species distribution
Sunburst controls
HideWeight segments by...
Change the size of the sunburst
Colour assignments
 Archea
Archea
|
 Eukaryota
Eukaryota
|
 Bacteria
Bacteria
|
 Other sequences
Other sequences
|
 Viruses
Viruses
|
 Unclassified
Unclassified
|
 Viroids
Viroids
|
 Unclassified sequence
Unclassified sequence
|
Selections
Click on a node to select that node and its sub-tree.
Clear selection
This visualisation provides a simple graphical representation of the distribution of this family across species. You can find the original interactive tree in the adjacent tab. More...
Tree controls
HideThe tree shows the occurrence of this RNA across different species. More...
Loading...
Please note: for large trees this can take some time. While the tree is loading, you can safely switch away from this tab but if you browse away from the family page entirely, the tree will not be loaded.
Trees
This page displays the predicted phylogenetic tree for the alignment. More...
Note: You can also download the data file for the seed tree.
Motif matches
There are 3 motifs which match this family.
This section shows the Rfam motifs that match sequences within the seed alignment of this family. Users should be aware that the motifs are structural constructs and do not necessarily conform to taxonomic boundaries in the way that Rfam families do. More...
| Original order | Motif Accession | Motif Description | Number of Hits | Fraction of Hits | Sum of Bits | Image |
|---|---|---|---|---|---|---|
| 7 | RM00008 | GNRA tetraloop | 2 | 0.250 | 21.8 |
|
| 7 | RM00024 | T-loop | 3 | 0.375 | 30.0 |
|
| 7 | RM00030 | U-turn motif | 2 | 0.250 | 21.8 |
|
References
This section shows the database cross-references that we have for this Rfam family.
Literature references
-
Ng WC, Soto-Acosta R, Bradrick SS, Garcia-Blanco MA, Ooi EE Viruses. 2017; [Epub ahead of print] The 5' and 3' Untranslated Regions of the Flaviviral Genome. PUBMED:28587300
-
Gebhard LG, Filomatori CV, Gamarnik AV Viruses. 2011;3:1739-1756. Functional RNA elements in the dengue virus genome. PUBMED:21994804
-
Zeng M, Duan Y, Zhang W, Wang M, Jia R, Zhu D, Liu M, Zhao X, Yang Q, Wu Y, Zhang S, Liu Y, Zhang L, Yu Y, Chen S, Cheng A Front Microbiol. 2020;11:473. Universal RNA Secondary Structure Insight Into Mosquito-Borne Flavivirus (MBFV) cis-Acting RNA Biology. PUBMED:32292394
-
Filomatori CV, Lodeiro MF, Alvarez DE, Samsa MM, Pietrasanta L, Gamarnik AV Genes Dev. 2006;20:2238-2249. A 5' RNA element promotes dengue virus RNA synthesis on a circular genome. PUBMED:16882970
-
Yu L, Nomaguchi M, Padmanabhan R, Markoff L Virology. 2008;374:170-185. Specific requirements for elements of the 5' and 3' terminal regions in flavivirus RNA synthesis and viral replication. PUBMED:18234265
-
Filomatori CV, Iglesias NG, Villordo SM, Alvarez DE, Gamarnik AV J Biol Chem. 2011;286:6929-6939. RNA sequences and structures required for the recruitment and activity of the dengue virus polymerase. PUBMED:21183683
-
Alvarez DE, Lodeiro MF, Luduena SJ, Pietrasanta LI, Gamarnik AV J Virol. 2005;79:6631-6643. Long-range RNA-RNA interactions circularize the dengue virus genome. PUBMED:15890901
-
Clyde K, Harris E; J Virol. 2006;80:2170-2182. RNA secondary structure in the coding region of dengue virus type 2 directs translation start codon selection and is required for viral replication. PUBMED:16474125
External database links
| Gene Ontology: | GO:0039694 (viral RNA genome replication); |
| Sequence Ontology: | SO:0000837 (UTR_region); |
Curation and family details
This section shows the detailed information about the Rfam family. We're happy to receive updated or improved alignments for new or existing families. Submit your new alignment and we'll take a look.
Curation
| Seed source | Lamkiewicz K | ||||||
| Structure source | Predicted; LocARNA 2.0.0RC8 | ||||||
| Type | Cis-reg; | ||||||
| Author |
Lamkiewicz K
|
||||||
| Alignment details |
|
Model information
| Build commands |
cmbuild -F CM SEED
cmcalibrate --mpi CM
|
| Search command |
cmsearch --cpu 4 --verbose --nohmmonly -T 30.00 -Z 2958934 CM SEQDB
|
| Gathering cutoff | 80.0 |
| Trusted cutoff | 82.4 |
| Noise cutoff | 62.1 |
| Covariance model | Download |
 Loading...
Loading...